当前位置:网站首页>[Space & single cellomics] phase 1: single cell binding space transcriptome research PDAC tumor microenvironment
[Space & single cellomics] phase 1: single cell binding space transcriptome research PDAC tumor microenvironment
2022-07-02 14:37:00 【Top bioinformation】
**
6.19On that day, I released the creation on gongzong 「 Space & Literature study group of single cell omics 」 's post , At the same time, several thinking problems have been put forward . Because the number of people is controlled , The final team members only identified9 position, There are more than ten friends who contact me , I missed the bus , To you 「 sorry 」. If there are other groups in the future , I will inform you again . Study groupcourse、videoAnd so on and so on 「 Gongzong No. was released 」, You can pay attention to it .
Last Sunday , The group conducted The first 1 period Share , from TOP bacteria and Jeffery be the speaker , Respectively introduced :
2020 Single cell binding spatial transcriptome study PDAC Tumor microenvironment science Recently published single cell space metabonomics technology
This tweet yes TOP Bacteria literature sharing Text version , video Explanation already in B standing Release ; The... Of this issue 2 Tweets yes Jeffery Literature sharing Text version , Reprinted from his gongzong No : Student information programming self-study room
Let's go to the literature interpretation
background
 This article was published in 2020 year 1 month , It is the earliest single cell binding idling article I can find .
This article was published in 2020 year 1 month , It is the earliest single cell binding idling article I can find .
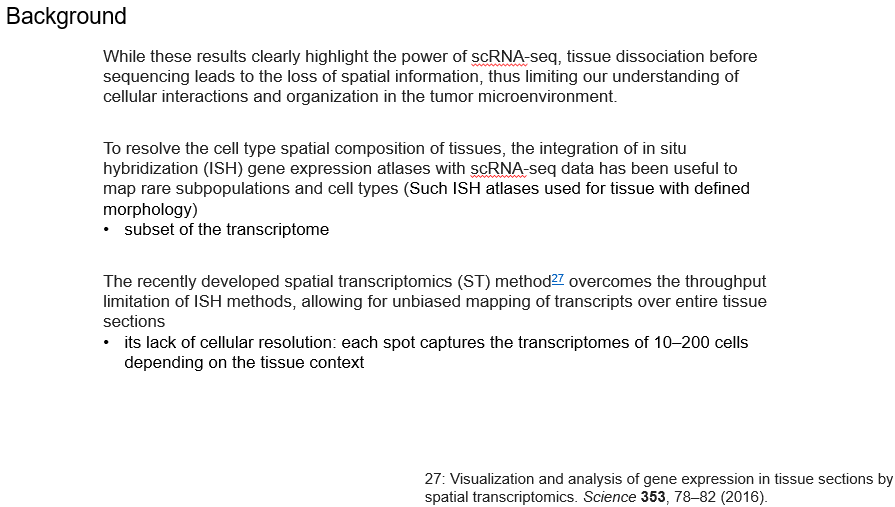 The background of the article is mainly 3 spot :
The background of the article is mainly 3 spot :
scRNA-seq It will be dissociated before sequencing , Loss of spatial information Combining in situ hybridization and scRNA-seq It can solve the above problems to a certain extent , but ISH The limitations are also obvious , Only a small number of genes can be captured As early as 2016 In the year , Space transcriptome technology was born , But the limitation is the lack of single cell resolution
*This article combines scRNA-seq And spatial transcriptome , The two technologies complement each other , To explore the tumor microenvironment of pancreatic ductal adenocarcinoma
*
result
1. Identify cell subpopulations
 The technical route is relatively simple , The sample size of early articles is also small , The main content is based on the samples of two patients ( Paired samples are scarce )
The technical route is relatively simple , The sample size of early articles is also small , The main content is based on the samples of two patients ( Paired samples are scarce )
 First, we made cluster annotation for single cell data of two patients , Then I took a look at A,B Consistency of annotation results of two patient subgroups .
First, we made cluster annotation for single cell data of two patients , Then I took a look at A,B Consistency of annotation results of two patient subgroups .
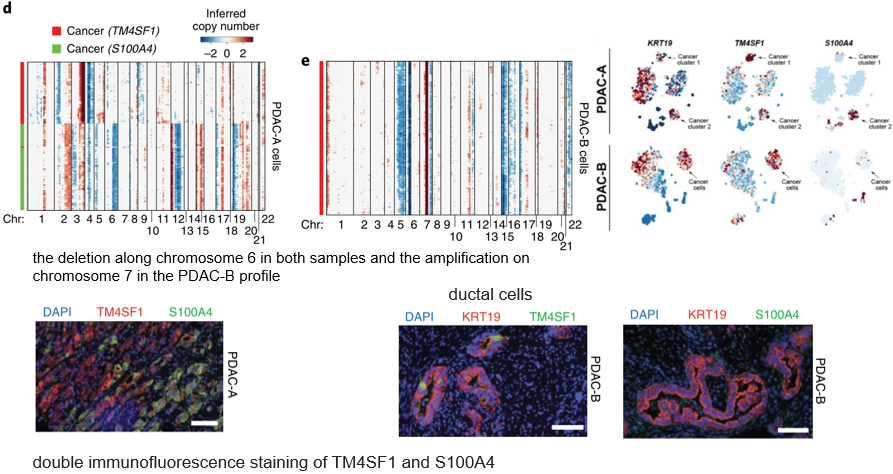 Still by inference CNV The malignant cells were identified by the method of , Further from CNV It can be seen from the heat map A There are two groups of patients CNV Cells with obvious differences . It was confirmed by immunofluorescence A,B The presence of tumor cells in the patient .
Still by inference CNV The malignant cells were identified by the method of , Further from CNV It can be seen from the heat map A There are two groups of patients CNV Cells with obvious differences . It was confirmed by immunofluorescence A,B The presence of tumor cells in the patient .
2. Analysis of spatial transcriptome data
 First of all, according to the HE Dyed pictures will A,B The patient's section is divided according to the tissue characteristics , From the spatial transcriptome data, we can also see some gene expression corresponding to the partition
First of all, according to the HE Dyed pictures will A,B The patient's section is divided according to the tissue characteristics , From the spatial transcriptome data, we can also see some gene expression corresponding to the partition
 Express data from idling only , Through the analysis process similar to single cell transcriptome , You can also see several groups spot, And it is consistent with the partition based on tissue slice
Express data from idling only , Through the analysis process similar to single cell transcriptome , You can also see several groups spot, And it is consistent with the partition based on tissue slice
3.MIA( Multimodal intersection analysis )
 In principle , Similar enrichment analysis , Is not complicated . The heat map on the right is viewed one by one , The deeper the red , It means that this cell type is in this region China and Vietnam are enriched . Through this analysis , Find out A Among patients , stay cancer region In addition to enrichment to tumor cells , It is also enriched in fibroblasts .
In principle , Similar enrichment analysis , Is not complicated . The heat map on the right is viewed one by one , The deeper the red , It means that this cell type is in this region China and Vietnam are enriched . Through this analysis , Find out A Among patients , stay cancer region In addition to enrichment to tumor cells , It is also enriched in fibroblasts .
4. Analysis of cell subclasses
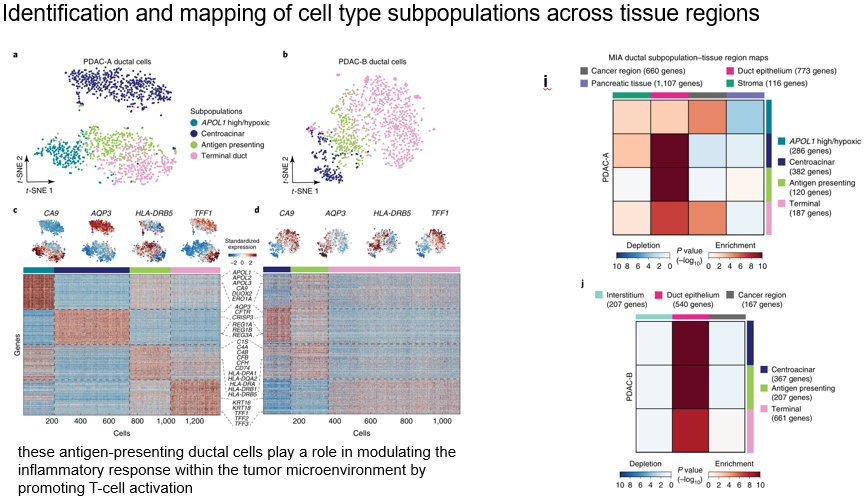
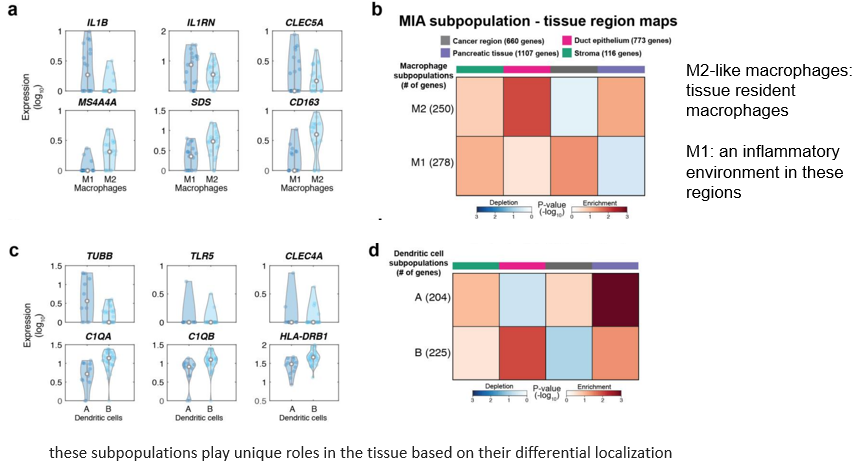 In this part, the ductal epithelial cells were analyzed in the same way 、 Macrophages 、DC cells . First, it is divided into sub categories , Reuse MIA Method to see the subclass in region The above enrichment .
In this part, the ductal epithelial cells were analyzed in the same way 、 Macrophages 、DC cells . First, it is divided into sub categories , Reuse MIA Method to see the subclass in region The above enrichment .
5. stay cancer region in , Whether different tumor cells co locate with different other cell types
 Take extra A Two sections of the patient , It is further subdivided in three slices cancer region. All three slices can be seen cancer cluster1 Co localization with fibroblasts .
Take extra A Two sections of the patient , It is further subdivided in three slices cancer region. All three slices can be seen cancer cluster1 Co localization with fibroblasts .
6. From the point of view of cell state scRNA-seq And idling
 utilize NMF From the single cell transcriptome data 3 Expression modules , The author focuses on stress-response Related modules .
utilize NMF From the single cell transcriptome data 3 Expression modules , The author focuses on stress-response Related modules . 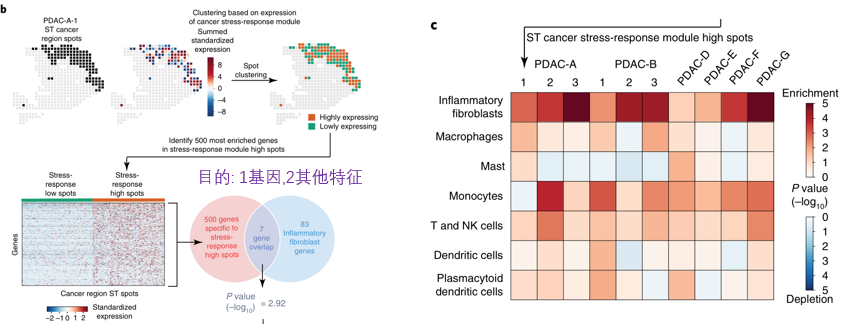 Then according to stress-response Module expression , take cancer region Of spot Divided into high and low groups , Continue the comparison between the two groups , find high This group of highly expressed genes . Obviously ,stress-response Related genes are already differential genes , Why do you do this ? There are at least two purposes for this :
Then according to stress-response Module expression , take cancer region Of spot Divided into high and low groups , Continue the comparison between the two groups , find high This group of highly expressed genes . Obviously ,stress-response Related genes are already differential genes , Why do you do this ? There are at least two purposes for this :
Increase the number of cells Reflect in addition stress-response Other features besides
The genes identified represent stress-response high Those of spot Regional characteristics of , And then use MIA Analysis stress-response high What cells will be enriched in the area of , Results show iCAF( Inflammatory fibroblasts ) Will be enriched in these areas .( There is a similar conclusion above , The conclusion here is to further refine .)
 TCGA Of bulk The sample also verified iCAF signature and stress-response module The relevance of
TCGA Of bulk The sample also verified iCAF signature and stress-response module The relevance of
This result was also confirmed by immunofluorescence .
The article also used mismatched melanoma samples , It goes further MIA Applicability of analysis .
summary

边栏推荐
- 微信小程序使用towxml显示公式
- php链表创建和遍历
- PHP linked list creation and traversal
- Methods of software testing
- Golang quickly generates model and queryset of database tables
- 篇9:XShell免费版安装
- Method of creating linked server for cross server data access
- 1. Editing weapon VIM
- Adhere to the foundation of 20 minutes go every day II
- Fabric. Keep the original level when JS element is selected
猜你喜欢

Onnx+tensorrt: write preprocessing operations to onnx and complete TRT deployment

测试框架TestNG的使用(二):testNG xml的使用

Certik released the defi security report in 2021, disclosing key data of industry development (PDF download link attached)

PyQt5_ Qscrollarea content is saved as a picture

一张图彻底掌握prototype、__proto__、constructor之前的关系(JS原型、原型链)

【空间&单细胞组学】第1期:单细胞结合空间转录组研究PDAC肿瘤微环境

STM32-DAC实验&高频DAC输出测试

YoloV6训练:训练自己数据集遇到的各种问题

Packet capturing tool Fiddler learning

Uniapp automated test learning
随机推荐
1、编辑利器vim
STM32库函数进行GPIO初始化
Fatal: unsafe repository is owned by someone else
STM32-DAC实验&高频DAC输出测试
taobao. trade. memo. Add (add remarks to a transaction) interface, Taobao store flag insertion interface, Taobao order flag insertion API interface, oauth2.0 interface
<口算練習機 方案開發原理圖>口算練習機/口算寶/兒童數學寶/兒童計算器 LCD液晶顯示驅動IC-VK1621B,提供技術支持
删除元素(带过渡动画)
一张图彻底掌握prototype、__proto__、constructor之前的关系(JS原型、原型链)
Fabric.js 橡皮擦的用法(包含恢复功能)
求轮廓最大内接圆
OpenHarmony笔记-----------(四)
What is erdma? Popular science cartoon illustration
Using computed in uni app solves the abnormal display of data () value in tab switching
Methods of software testing
由粒子加速器产生的反中子形成的白洞
STM32 standard firmware library function name memory (II)
docker mysql
MQ教程 | Exchange(交换机)
Data Lake (11): Iceberg table data organization and query
Fabric.js 元素被选中时保持原有层级