当前位置:网站首页>Bedtools tutorial
Bedtools tutorial
2022-07-02 11:36:00 【qq_ twenty-seven million three hundred and ninety thousand and 】
bedtools: a powerful toolset for genome arithmetic
bedtools Tools are a powerful tool for a wide range of genomic analysis tasks . The most widely used tool enables genome arithmetic : That is, the set theory on the genome . for example ,bedtools Allow people to start from the widely used genome file format ( Such as BAM、BED、GFF/GTF、VCF) Cross in multiple files of 、 Merge 、 Count 、 Complement and shuffle genome intervals . Although each individual tool is designed to do a relatively simple task ( for example , Intersect with two interval files ), But through in UNIX Combine multiple on the command line bedtools The operation can carry out quite complex analysis .
Genome annotation file download address
https://genome.ucsc.edu/cgi-bin/hgTables download bed file
bedtools --version # Version number
bedtools --contact # Help information
# Download the test file
curl -O https://s3.amazonaws.com/bedtools-tutorials/web/cpg.bed
curl -O https://s3.amazonaws.com/bedtools-tutorials/web/exons.bed
curl -O https://s3.amazonaws.com/bedtools-tutorials/web/gwas.bed
curl -O https://s3.amazonaws.com/bedtools-tutorials/web/genome.txt
###1 bedtools intersect
## Calculation overlap intervals
#Tool: bedtools intersect (aka intersectBed)
#Version: v2.30.0
#Summary: Report overlaps between two feature files.
#Usage: bedtools intersect [OPTIONS] -a <bed/gff/vcf/bam> -b <bed/gff/vcf/bam>
# notes :-b You can connect multiple files
# Show cpg.bed neutralization exons.bed There are overlapping intervals
bedtools intersect -a cpg.bed -b exons.bed
# Show exons.bed neutralization cpg.bed There are overlapping intervals
bedtools intersect -a exons.bed -b cpg.bed
# At the same time, the overlapping areas are displayed A、B Original records in the document
bedtools intersect -a exons.bed -b cpg.bed -wa -wb
# Show the base number of overlapping areas
bedtools intersect -a cpg.bed -b exons.bed -wo
# Show each one cpg.bed The record in the file is exons.bed Number of overlapping records in the file
bedtools intersect -a cpg.bed -b exons.bed -c
# cpg.bed There is a disagreement in the document exons.bed whatever intervals Overlapping records
bedtools intersect -a cpg.bed -b exons.bed -v
bedtools intersect -a cpg.bed -b exons.bed -wo
# Set threshold , Show cpg.bed in intervals There are at least 50% Sequence sum exons.bed Overlap in
bedtools intersect -a cpg.bed -b exons.bed -wo -f 0.50
# Overlapping areas of multiple files
bedtools intersect -a cpg.bed -b gwas.bed exons.bed
bedtools intersect -a cpg.bed -b gwas.bed exons.bed -wa -wb -names gwas exon # Add files label
# sorted Data is added -sorted Parameters , Run faster
time bedtools intersect -a exons.bed -b cpg.bed gwas.bed -sorted >>/dev/null
###2 bedtools merge
#Tool: bedtools merge (aka mergeBed)
#Version: v2.30.0
#Summary: Merges overlapping BED/GFF/VCF entries into a single interval.
#Usage: bedtools merge [OPTIONS] -i <bed/gff/vcf>
# Be careful :bedtools merge The input file is required to be sorted first
# Sort , Input files are sorted by chromosome first , Then sort by starting position .
sort -k1,1 -k2,2n test.bed >test.sorted.bed
# Show the final " Merge " Section
bedtools merge -i exons.bed | head -n 20
# The calculation leads to every new " Merge " The number of overlapping intervals , We will " Calculation " First column .
bedtools merge -i exons.bed -c 1 -o count | head -n 20
# Show all merged into new " Merge " The second row of the overlapping interval
bedtools merge -i exons.bed -c 2 -o collapse | head -n 20
# Merge distance does not exceed 1000 The range of ,
bedtools merge -i exons.bed -d 1000 -c 1 -o count | head -20
# Merge distance does not exceed 90 Area , Do different operations on the first column and the fourth column respectively
bedtools merge -i exons.bed -d 90 -c 1,4 -o count,collapse | head -20
###3 bedtools complement
#Tool: bedtools complement (aka complementBed)
#Version: v2.30.0
#Summary: Returns the base pair complement of a feature file.
#Usage: bedtools complement [OPTIONS] -i <bed/gff/vcf> -g <genome>
# notes :The genome file should tab delimited and structured as follows:
# <chromName><TAB><chromSize>
# genome.txt in ,exons.bed There is no interval
bedtools complement -i exons.bed -g genome.txt
###4 bedtools genomecov
#Tool: bedtools genomecov (aka genomeCoverageBed)
#Version: v2.30.0
#Summary: Compute the coverage of a feature file among a genome.
#Usage: bedtools genomecov [OPTIONS] -i <bed/gff/vcf> -g <genome>
# notes : Need sorted files
bedtools genomecov -i exons.bed -g genome.txt
# Output BEDGRAPH, Calculation intervals Of depth
bedtools genomecov -i exons.bed -g genome.txt -bg | head -20
###5 bedtools jaccard
#Tool: bedtools jaccard (aka jaccard)
#Version: v2.30.0
#Summary: Calculate Jaccard statistic b/w two feature files.
# Jaccard is the length of the intersection over the union.
# Values range from 0 (no intersection) to 1 (self intersection).
#Usage: bedtools jaccard [OPTIONS] -a <bed/gff/vcf> -b <bed/gff/vcf>
# Calculate similarity
bedtools jaccard -a cpg.bed -b exons.bed
###6 bedtools coverage
#Tool: bedtools coverage (aka coverageBed)
#Version: v2.30.0
#Summary: Returns the depth and breadth of coverage of features from B
# on the intervals in A.
#Usage: bedtools coverage [OPTIONS] -a <bed/gff/vcf> -b <bed/gff/vcf>
bedtools coverage -a cpg.bed -b exons.bed
Reference resources :
http://quinlanlab.org/tutorials/bedtools/bedtools.html
边栏推荐
- PHP tea sales and shopping online store
- Tick Data and Resampling
- 八大排序汇总
- ctf 记录
- TIPC Cluster5
- The working day of the month is calculated from the 1st day of each month
- JS——每次调用从数组里面随机取一个数,且不能与上一次为同一个
- 2022年遭“挤爆”的三款透明LED显示屏
- TIPC Service and Topology Tracking4
- Is the stock account given by qiniu business school safe? Can I open an account?
猜你喜欢
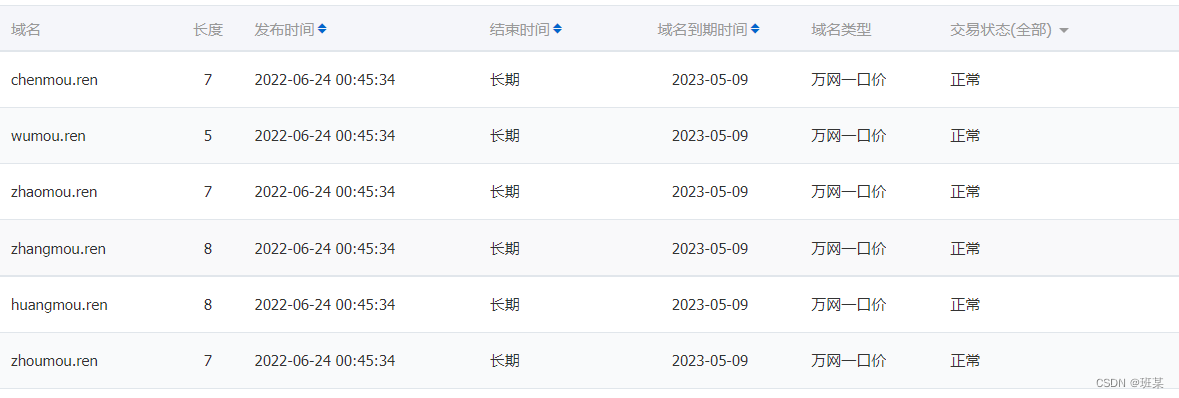
Is the Ren domain name valuable? Is it worth investing? What is the application scope of Ren domain name?
![[cloud native] 2.5 kubernetes core practice (Part 2)](/img/87/826894d758392a0c7a60dd5fa09eef.png)
[cloud native] 2.5 kubernetes core practice (Part 2)
Mmrotate rotation target detection framework usage record
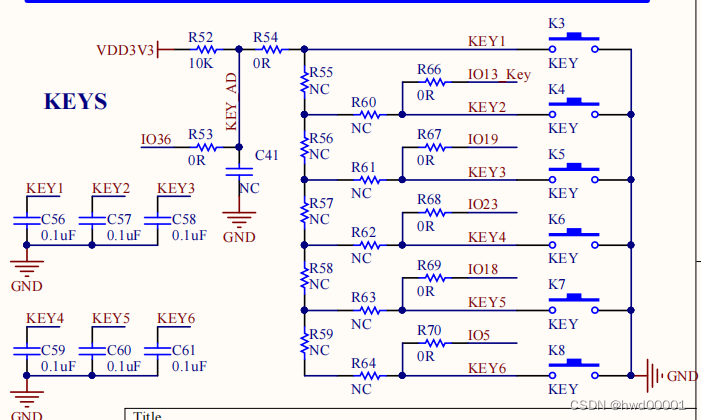
ESP32音频框架 ESP-ADF 添加按键外设流程代码跟踪
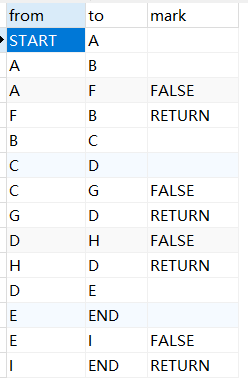
mysql链表数据存储查询排序问题

Principle of scalable contract delegatecall
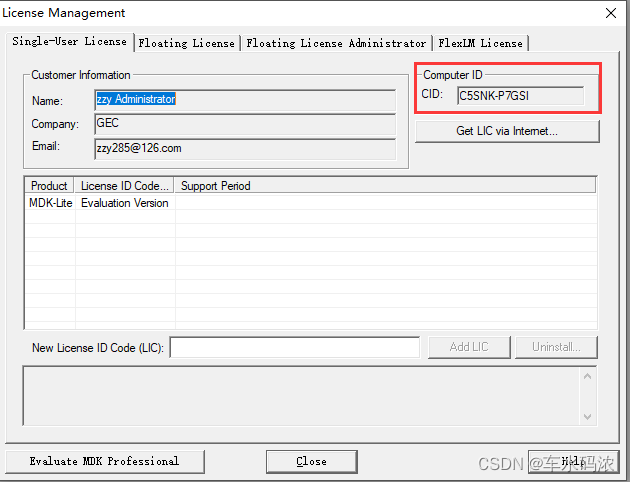
I STM32 development environment, keil5/mdk5.14 installation tutorial (with download link)
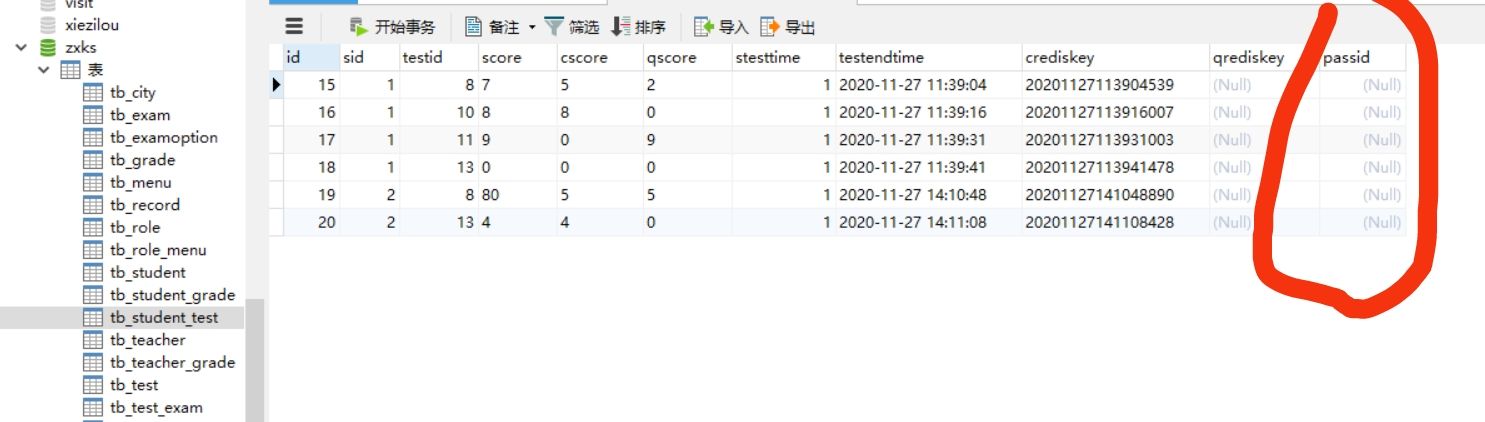
Map set assignment to database
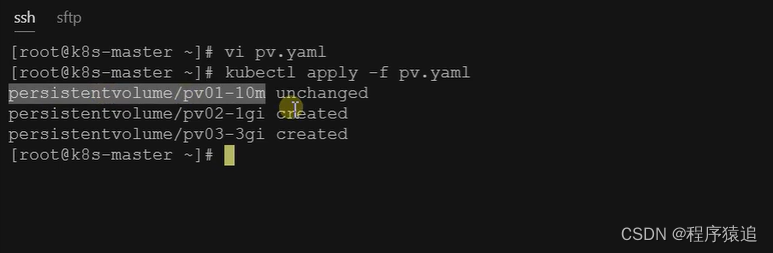
【云原生】2.5 Kubernetes 核心实战(下)
基于Hardhat和Openzeppelin开发可升级合约(一)
随机推荐
二.Stm32f407芯片GPIO编程,寄存器操作,库函数操作和位段操作
sqlite 修改列类型
Programmer growth Chapter 6: how to choose a company?
tqdm的多行显示与单行显示
Is the Ren domain name valuable? Is it worth investing? What is the application scope of Ren domain name?
ESP32音频框架 ESP-ADF 添加按键外设流程代码跟踪
Gaode draws lines according to the track
解决uniapp列表快速滑动页面数据空白问题
Thanos Receiver
C#基于当前时间,获取唯一识别号(ID)的方法
III Chip startup and clock system
SSRF
揭露数据不一致的利器 —— 实时核对系统
启牛商学院给的股票账户安全吗?能开户吗?
Tick Data and Resampling
Win11 arm system configuration Net core environment variable
ros缺少catkin_pkg
Some things configured from ros1 to ros2
可升级合约的原理-DelegateCall
STM32 single chip microcomputer programming learning